High performance computing enables real-time 3D visualisations of embryo development, allowing scientists to better understand the "emergence of life" from cells
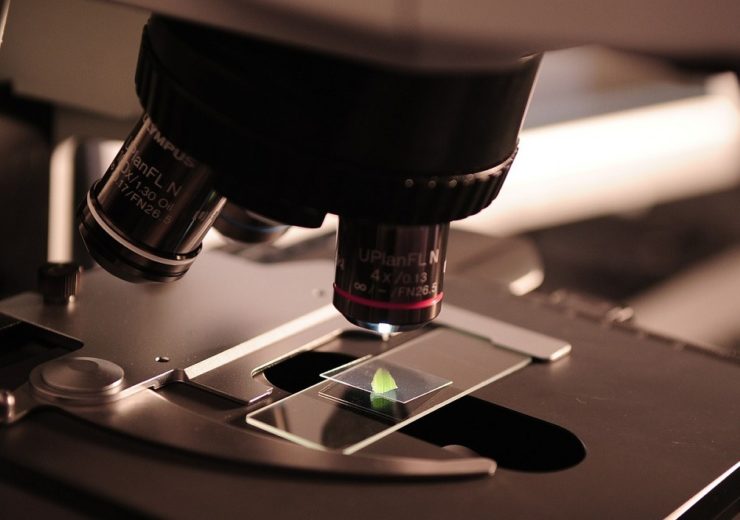

Scientists are using supercomputers to observe single cells with high precision
High-powered supercomputers are being used by scientists to simulate embryological development in a bid to better understand the building blocks of life.
By harnessing the processing power of these high performance computing (HPC) machines, biologists can observe the development cycle of single cells in virtual reality environments, and use these to examine how complex organisms are formed.
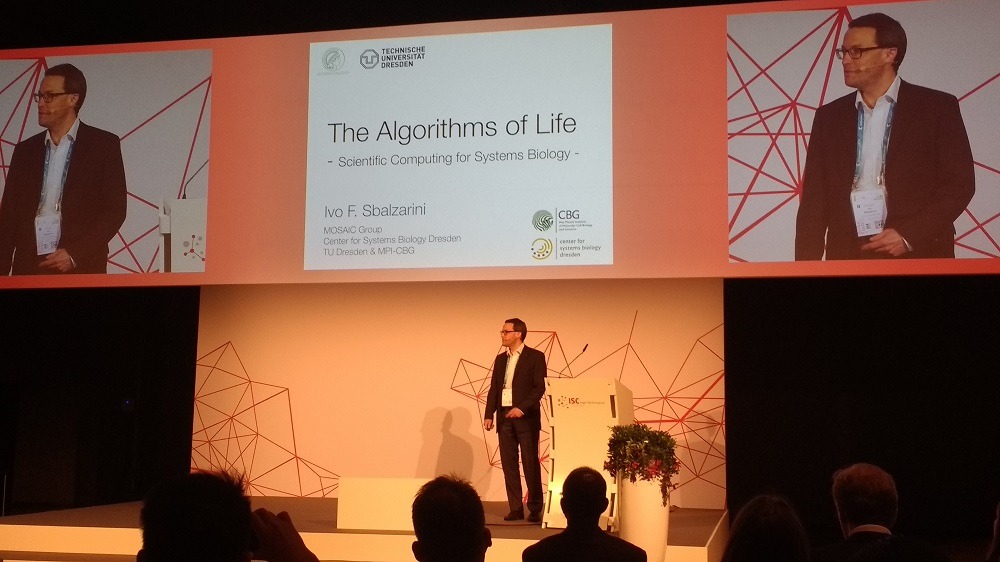
In a keynote address at the ISC High Performance conference in Frankfurt yesterday, Professor Ivo Sbalzarini – who holds senior positions at Technische Universität Dresden and the Max Planck Institute of Molecular Cell Biology and Genetics – explained that researchers like himself are now actively studying the “very emergence of life from a single cell”.
Observing embryo development in this way is hoped to provide “new explanations for diseases” and even develop new forms of treatment.
Mr Sbalzarini also revealed interesting similarities between the design and activity of both the cells he studies and the computers being used to digitally replicate them.
Supercomputers use combined processing power to solve complex problems
Supercomputers such as IBM’s Summit and Sierra, the world’s two fastest computing machines, work by aggregating the processing power of multiple computers into “clusters”, which operate collectively to dramatically increase the potential of each individual unit – or “node”.
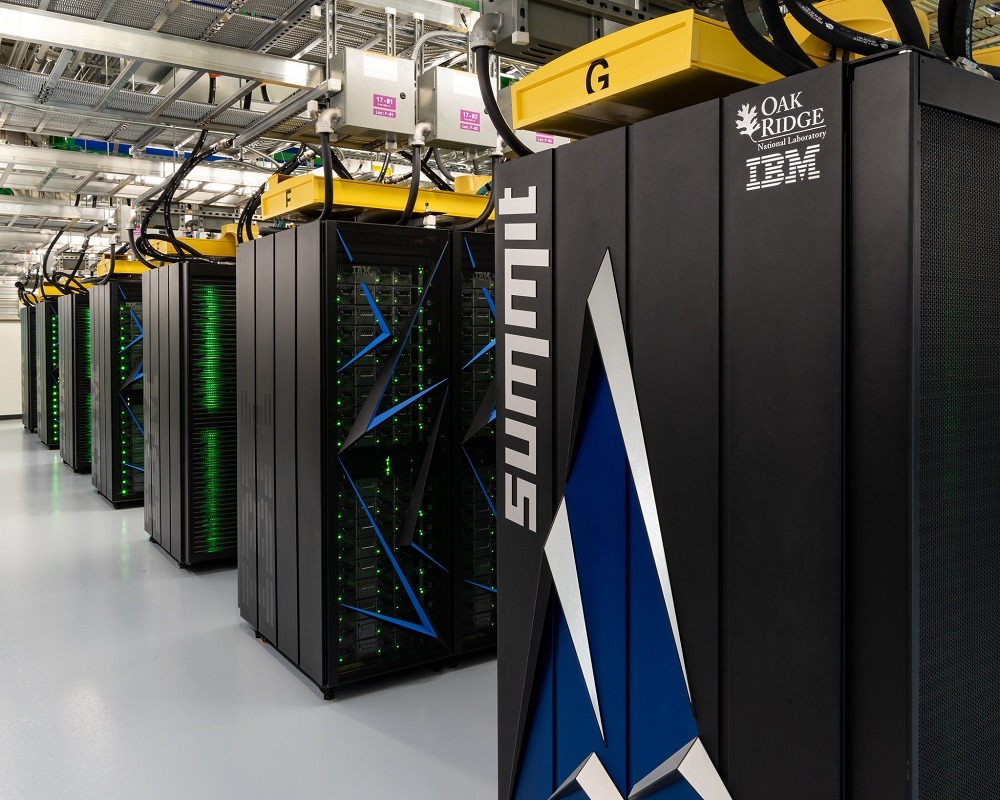

Typical top-level HPC machines combine thousands of processors, each of which operates individually as well as being part of the wider network.
They demand enormous – and expensive – amounts of upkeep in terms of power, cooling and other maintenance, but when operating at capacity they are able to deliver the vast computational power needed to solve complex mathematical problems.
To simulate the development of an organism even as simple as a fruit fly, which Mr Sbalzarini gave as a demonstration of his work, the processing power of HPC is necessary.
Mr Sbalzarini said: “The first thing you want to do is look at these images, which might sound like something simple but it’s made hard by the sheer size of the data sets.
“We have high resolution 3D microscopes that can image embryos over the lifespan of development, in order to study tissue formation and the very emergence of life from a single cell.
“The image is decomposed into blocks which are sent to different processing nodes, so one node only really has to solve a very small problem.
“Collectively the processors solve the global problem, and they do it so fast that we are able to see it in real time.”
Embryological development can be viewed in real-time virtual reality
By using supercomputers, scientists are able to take the images they see through the microscope and create a virtual copy which can be observed and mapped more clearly than previously possible.
Mr Sbalzarini added: “When we image the fruit fly embryo over the 72 hours of development this is about 180 terabytes of image data.

“We need a rendering performance of about 1.8 gigapixels per second to directly view the images from the microscope as they are being acquired in a virtual reality or augmented reality environment.
“This allows the scientist to immerse themselves in the developing embryo, to walk around it and stick their head inside to see what the developing cells are doing on the inside – not just the outside as is typically the case with a microscope.
“The computer identifies the location and shape of every cell in the tissue in order to track it and describe it.
“The motivation is to understand biology and understand how cells form tissues and to provide new explanations for diseases and maybe other therapies for disease.”
High performance computing has similarities to embryological development
Simulating embryological development requires enormous computer power, the complexity of which depends heavily on the collective processing power of HPC technology.
Mr Sbalzarini drew comparisons between the supercomputers that make his work possible and the very structure of the organisms he studies.
He said: “There is no master cell – no boss cell – that can tell the others what to do at any point in time and space.
“Instead, what we are looking at is a massively parallel and fully self-organised system in which we can view every single cell as a processing element that executes programmes.
“The language in biology already makes that clear when we talk about genetic programmes, cellular decision-making and signalling networks.
“The link to computer science is clear as we talk about programmes, networks and sorting.
“Cells do not do that in isolation, they are connected to each other and they communicate via a multitude of signals – chemical, hormone and mechanical.
“For this massively parallel computer, if every cell is the processing element, we are talking about a highly interconnected computing system here, which is able to break several complexity limits and with billions of processing elements.”